Jae Il Lyu1![]() , Chaewon Lee2
, Chaewon Lee2![]() , Minwoo Kim3
, Minwoo Kim3![]() , HwangWeon Jeong3
, HwangWeon Jeong3![]() , Song Lim Kim1
, Song Lim Kim1![]() , JeongHo Baek1
, JeongHo Baek1![]() , Kyung-Hwan Kim3,*
, Kyung-Hwan Kim3,*![]()
1Digital Breeding Convergence Division, National Institute of Agricultural Science, RDA, Jeonju 54874, Republic of Korea
2Crop Production and Physiology Research Division, National Institute of Crop Science, RDA, Jeonju 55365, Republic of Korea
3Supercomputing Center, National Institute of Agricultural Science, RDA, Jeonju 54874, Republic of Korea
Correspondence to Kyung-Hwan Kim, E-mail: biopiakim@Korea.kr
Plant Image Sci. 1:4. https://doi.org/10.65971/PIS.2025.1.4
Received on November 06, 2025, Revised on December 05, 2025 , Accepted on December 17, 2025 , Published on December 31, 2025.
© Author(s). This is an Open Access article distributed under the terms of the Creative Commons CC BY 4.0 (https://creativecommons.org/licenses/by/4.0/) which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Global demographic expansion and accelerating climate change are heightening the need for sustainable enhancement of crop productivity and resilience to environmental stresses. As conventional breeding approaches based on empirical selection approach their limits, digital breeding is emerging as an integrated framework that combines high-throughput imaging, multi-source data, and artificial intelligence (AI). Although next-generation sequencing (NGS) and smart-farming systems have enabled large-scale accumulation of genomic and environmental datasets, the phenotypic dimension remains a critical bottleneck in predictive breeding. To overcome this limitation, Korea has established six large-scale national phenotyping platforms equipped with advanced RGB, hyperspectral, and thermal imaging systems for high-throughput, precision phenotypic data acquisition. These platforms support quantitative and non-destructive monitoring of plant morphology, stress responses, and developmental dynamics and employ AI-driven classification and predictive modeling to derive biologically relevant traits. The integration of genomic, phenomic, and environmental datasets through AI-based analytical pipelines is expected to accelerate digital breeding, facilitating the development of climate-resilient and consumer-oriented cultivars within a data-driven agricultural paradigm. Collectively, plant phenomics research is evolving beyond image-based observation toward a comprehensive, predictive framework that provides the technological basis for next-generation precision agriculture.
artificial intelligence, digital breeding, high-throughput imaging, plant phenomics
The global population, currently estimated at 7.5 billion, is projected to reach nearly 9 billion by 2050 (Gerland et al. 2014), presenting one of the greatest challenges of the twenty-first century—securing sufficient food and fuel resources to sustain humanity. To meet the increasing demand for food, crop productivity must increase by approximately 38% per year that necessitates innovative breeding programs that surpass the limitations of traditional experience-based approaches (Baker 1986). Conventional breeding systems, that are traditionally dependent on human intuition, are increasingly being transformed through digital breeding that integrates imaging technologies, large-scale data analytics, and artificial intelligence (AI) to facilitate environmentally contextualized decision-making in crop selection. Digital breeding converts genomic, phenotypic, and environmental information into digital formats by leveraging big data and AI-driven analytics to identify and predict traits that address ecological sustainability and consumer preferences (Jeon et al. 2023).
To operationalize this digital breeding framework, comprehensive datasets encompassing the genomic, environmental, and phenotypic dimensions are indispensable. Although genomic and environmental datasets have proliferated with advances in next-generation sequencing (NGS) and smart farming technologies, phenotypic information remains comparatively scarce. Traditional phenotyping is labor-intensive and time-consuming, whereas digital phenotyping enables the rapid high-throughput quantification of traits—including those undetectable by conventional observation—using advanced imaging sensors and computational analysis (Furbank et al. 2019). Digital phenotyping systems employ red–green–blue (RGB), near-infrared (NIR), fluorescence, and hyperspectral imaging to quantitatively assess plant traits such as morphology and canopy architecture and translate visual signals into numerical data that are quantitatively associated with agronomic performance indicators such as growth rate, yield, and stress tolerance. Phenomics is a multidisciplinary science that investigates plant traits resulting from genotype–environment interactions, encompassing morphological, physiological, metabolic, and stress-response parameters (Deery and Jones 2021). Thus, the integration of genomic, phenomic, and environmental data forms the foundation for accelerating the development of climate-resilient and consumer-oriented cultivars. The following sections outline the current landscape of phenomic infrastructures and research trends in Korea that facilitate the implementation of digital breeding.
Over the past decade, Korea has developed a cohesive national network of phenomic research facilities, covering both controlled and field conditions. Six major institutions operate large-scale phenotyping infrastructures equipped with advanced imaging sensors, encompassing visible, infrared, fluorescence, and hyperspectral systems (Fig. 1).
Early-stage systems relied on imported platforms such as LemnaTec and PSI; however, recent advances spearheaded by the Korea Institute of Science and Technology (KIST) have facilitated the development of domestically engineered, fully integrated phenotyping platforms. The National Institute of Agricultural Sciences (NAS) manages a comprehensive phenomics complex with high-throughput imaging systems, precision measurement laboratories, and smart greenhouses. Since 2017, it has conducted large-scale analyses of rice and soybean focusing on drought tolerance, seed morphology, and image-based disease assessments using automated pipelines. The National Institute of Crop Science (NICS) operates semi-controlled greenhouses equipped with robotic XYZ imaging systems for herbicide damage diagnosis, drought index development, and predictive modeling of crop growth, whereas outdoor field scanners monitor the rice canopy dynamics. The National Institute of Horticultural and Herbal Science (NIHHS) uses Venlo-type and conveyor-based greenhouses and environmental control chambers to study horticultural crop physiology, stress tolerance, and spectral index development. The KIST operates climate-controlled chambers and conveyor systems for chlorophyll fluorescence and 3D phenotyping research, integrating genomic, metabolomic, and environmental data to identify functional traits. The Korea Atomic Energy Research Institute (KAERI) applies imaging diagnostics and radiation mutation analyses to develop AI-assisted preselection models and digital growth indicators. The Daegu Gyeongbuk Institute of Science and Technology (DGIST) maintains phenotyping facilities for Arabidopsis thaliana, focusing on life-cycle phenotyping, leaf senescence quantification, and abiotic stress response analysis. Universities have contributed to stress quantification, seed characterization, and algorithm development through targeted studies using RGB, IR, and hyperspectral imaging. An effective phenomics implementation requires the integration of imaging, automation, computation, and control technologies. Despite the limited domestic market, Korean companies, such as Phenobox, have contributed to system integration and strengthened national capacity in smart agriculture. Over the past decade, Korean phenomics research has rapidly progressed from limited resources to an organized national network. The Korean Phenomics Research Society, founded in 2015, evolved into the Korean Society of Plant Image Science in 2024, connecting more than 500 researchers through annual conferences and workshops and fostering collaboration and innovation in digital plant phenotyping.
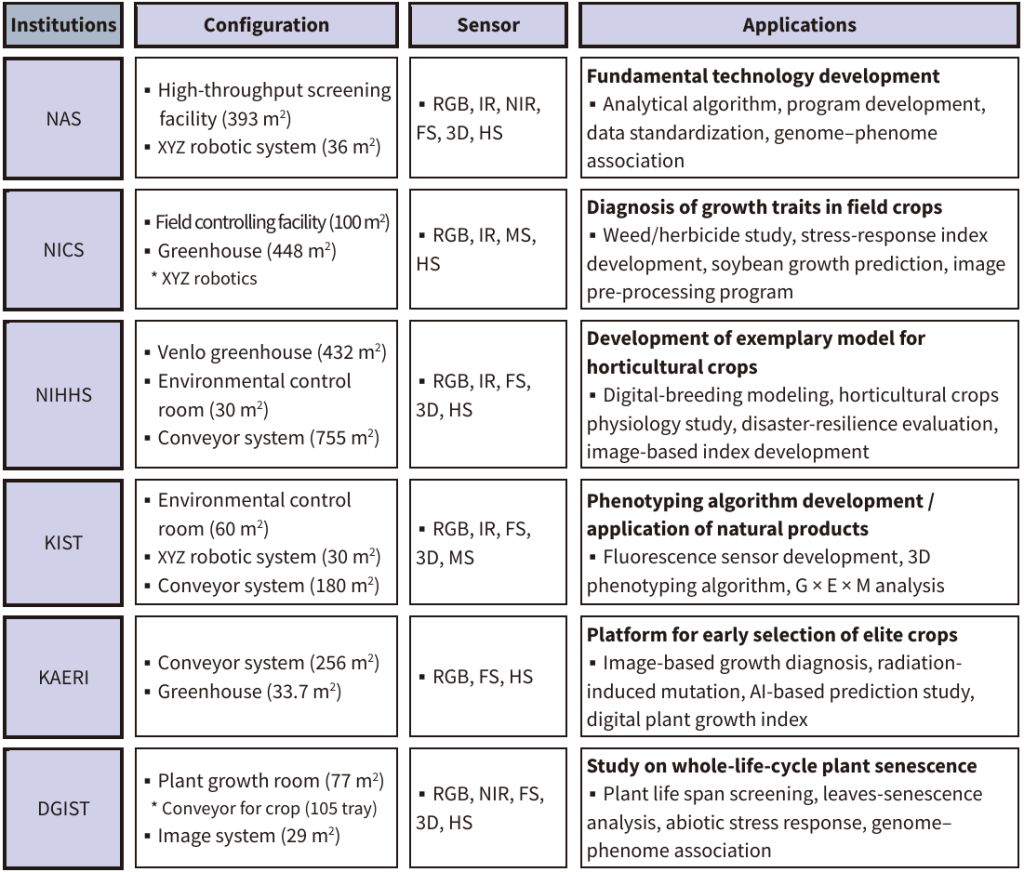
Fig. 1. Status of domestic high-throughput phenomics facilities and their research applications. NAS, National Academy of Agricultural Sciences; NICS, National Institute of Crop Science; NIHHS, National Institute of Horticultural and Herbal Science; KIST, Korea Institute of Science and Technology; KAERI, Korea Atomic Energy Research Institute; DGIST, Daegu Gyeongbuk Institute of Science and Technology; RGB, visible; IR, infrared; NIR, near-infrared; FS, fluorescence; 3D, three dimensions; HS, hyperspectral; MS, multispectral; G × E × M, genotype × environment × management; AI, artificial intelligence.
A recent meta-analysis of 67 Korean studies published over the past five years identified five major research directions in plant phenomics. (1) Early investigations primarily examined morphological traits such as leaf area and stem architecture; however, the research scope has since broadened to encompass seed and fruit traits, abiotic stress responses, and disease diagnostics. (2) Target crops have shifted from model species to economically important staples such as rice, soybean, wheat, barley, and maize, with a heightened focus on drought, salinity tolerance, and photosynthetic efficiency. (3) Imaging technologies have evolved from simple RGB-based systems to thermal, fluorescence, LiDAR, 3D, and hyperspectral modalities, allowing for the detailed physiological characterization of parameters such as water status, leaf temperature, and photosynthetic activity. (4) Currently, AI plays a pivotal role in phenomic data interpretation, employing CNN, U-Net, Mask R-CNN, Transformer, and Segment Anything Model (SAM) architectures for classification, segmentation, and predictive modeling with a high predictive accuracy. (5) Integrative modeling has merged phenotypic, genotypic, and environmental datasets to improve predictive capacity and inform breeding decision-making. Recent advances in digital twin and AI-based soft-sensor frameworks have facilitated real-time monitoring and adaptive control in smart farming environments. (Table 1).
Table 1. Overview of domestic phenomics research and associated sensor applications
| Trait | Plant | Sensor | Key content | Research (No.) |
|---|---|---|---|---|
| Morphological phenotyping | Rice, soybean, tomato, Arabidopsis | RGB, infrared, hyperspectral, near-infrared | – Imaging of non-specific physiological traits : Growth rate, moisture content, biomass, disease – Morphological analysis via U-Net and Mask R-CNN – Quantification of leaf area and stem structure using 3D reconstruction | 18 |
| Stress tolerance | Rice, soybean, Arabidopsis | RGB, hyperspectral, near-infrared | – Quantification of physiological and photosynthetic responses to drought, salinity, and temperature stress – Early diagnosis using hyperspectral and infrared data | 13 |
| Seed/grain | Rice, soybean, tomato | RGB, near-infrared | – Automatic analysis of grain development and quality – Prediction of protein and starch content via hyperspectral imaging | 15 |
| Root | Rice, soybean | 3D scanner, X-ray, microCT | – 3D root structure analysis with quantification of length and volume – Root segmentation, GWAS-based association analysis | 17 |
| AI classification | Wheat, tomato | RGB, hyperspectral | – AI-based classification of crop diseases, morphology, and quality – Application of transformer and ViT models – Enhancement of prediction accuracy | 14 |
RGB, red-green-blue; R-CNN, region-based convolutional neural network; 3D, three dimensions; microCT, micro-computed tomography; GWAS, genome-wide association study; AI, artificial intelligence; ViT, vision transformer.
Practical applications involve the longitudinal imaging of drought-tolerant transgenic rice using RGB and NIR sensors, combined with soil moisture measurements to visualize drought-rehydration dynamics (Kim et al. 2020). Automated irrigation systems such as DroughtSpotter have been adopted for the quantitative assessment of evapotranspiration and water-use efficiency under controlled high-temperature and low-humidity conditions. In seed phenotyping, RGB imaging of 39,065 soybean seeds across 400 accessions has enabled the extraction of eight morphological parameters—area, perimeter, width, height, thickness, circularity, roundness, and solidity—as well as color indices that has facilitated variety classification and GWAS integration (Baek et al. 2020). Similar analytical pipelines have been extended to other crops including mung beans, sprouts, and ornamental plants. Furthermore, mobile field phenotyping platforms developed by the NAS and NIHHS have been deployed for canopy-level imaging aimed at yield prediction and early disease detection in paddy and orchard systems. The National Information Society Agency (NIA) has released more than 40 agricultural AI datasets through its AI-Hub platform, covering phenotypic, environmental, and smart farm data for multiple crops, thereby promoting the application of AI models in agricultural research. In 2023, the NAS Crop PhenomicsCenter was designated as the national reference standard for phenotypic data, registering longitudinal datasets of rice and soybeans to establish digital benchmarks and ensure future data interoperability and reusability.
The growing global population and intensifying climate challenges have prompted a paradigm shift in crop breeding—from traditional empirical approaches to digital breeding—supported by high-throughput phenotyping and AI-driven analytics. Over the past decade in Korea, sustained investment in phenomics research has led to substantial progress in imaging technology, data integration, and model development. Several major trends have emerged: research has expanded from structural analyses of leaves and roots to include seeds, fruits, stress tolerance, and disease detection; focus has shifted from model species to major crops such as rice, soybeans, wheat, barley, and maize; imaging technologies have evolved from RGB-based systems to multimodal platforms incorporating thermal, fluorescence, LiDAR, and hyperspectral sensing; AI-based analytical models—such as CNN, U-Net, Mask R-CNN, Transformer, and SAM— have demonstrated high predictive accuracy in detection, classification, and growth prediction; finally, integrative approaches linking phenotypic, genotypic, and environmental data have facilitated comprehensive modeling of plant physiology and adaptive responses. These efforts have resulted in the development of digital twin and AI-based soft-sensor frameworks that facilitate real-time monitoring and model-based control in smart agricultural systems. Collectively, plant phenomics is evolving into a data-driven agricultural discipline that integrates AI, multi-sensor imaging, and genomics. This convergence lays the foundation for predictive and precision agriculture, supporting accurate yield estimation, stress diagnostics, and automated crop management. As digital twin systems and real-time monitoring technologies continue to advance, the realization of fully autonomous, data-centric smart farming is expected to reshape the paradigm of sustainable agriculture in the coming decade.
The authors thank Hongseok Lee (NICS), Woo-Moon Lee (NIHHS), Hyung-Seok Kim (KIST), and Jin-Baek Kim (KAERI) for their assistance with infrastructure information.
Conceptualization: Baek JH, Kim KH; Investigation and analysis: Lee C, Kim M, Jeong HW; Writing – original draft: Lyu JI, Kim KH; Writing – review and editing: Lee C, Kim M, Jeong HW, Kim SL, Baek JH, Kim KH.
The authors declare no conflicts of interest.
This study was supported by the “Cooperative Research Program for Agriculture Science and Technology Development (Project No. RS-2021-RD010130)” of the Rural Development Administration, Republic of Korea.
The data supporting the findings of this study are available from the corresponding author upon reasonable request.
Baek JH et al. 2020. High throughput phenotyping for various traits on soybean seeds using image analysis. Sensors. 20(1):248. https://doi.org/10.3390/s20010248
Baker RJ. 1986. Selection indices in plant breeding. Boca Raton (FL): CRC Press. https://doi.org/10.1201/9780429280498
Deery DM, Jones HG. 2021. Field phenomics: will it enable crop improvement? Plant Phenomics. 2021:9871989. https://doi.org/10.34133/2021/9871989
Furbank RT, Jimenez-Berni JA, George-Jaeggli B, Potgieter AB, Deery DM. 2019. Field crop phenomics: enabling breeding for radiation use efficiency and biomass in cereal crops. New Phytol. 223(4):1714–1727. https://doi.org/10.1111/nph.15817
Gerland P et al. 2014. World population stabilization unlikely this century. Science. 346(6206):234–237. https://doi.org/10.1126/science.1257469
Jeon D et al. 2023. Digitalizing breeding in plants: a new trend of next-generation breeding based on genomic prediction. Front Plant Sci. 14:1092584. https://doi.org/10.3389/fpls.2023.1092584
Kim SL et al. 2020. High-throughput phenotyping platform for analyzing drought tolerance in rice. Planta. 252:38. https://doi.org/10.1007/s00425-020-03436-9